Friedman Test
- Home /
- Friedman
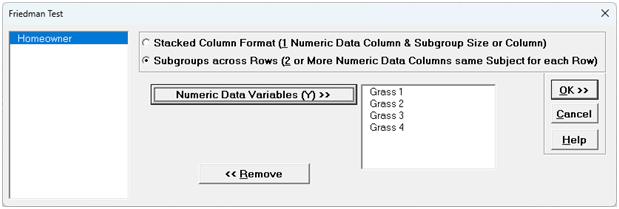
The Friedman test is an extension of the Wilcoxon Signed-Rank test and the nonparametric equivalent of a One-Way Repeated-Measures ANOVA(i.e., One-Way ANOVA on Within Subject Ranks). The concept is the same as that of the Paired t-test, we are interested in the within subject variability, not the subject-to-subject variability.
It can also be considered as a nonparametric alternative to a DOE model with blocks or GLM with one fixed factor and one random factor. The Friedman test requires exactly 1 non-missing observation for each level of the treatment, within each block.
The null and alternative hypothesis are:
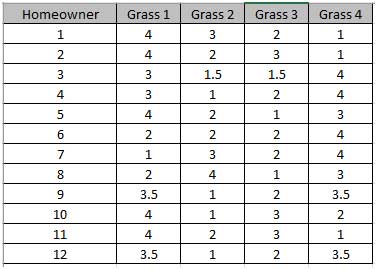
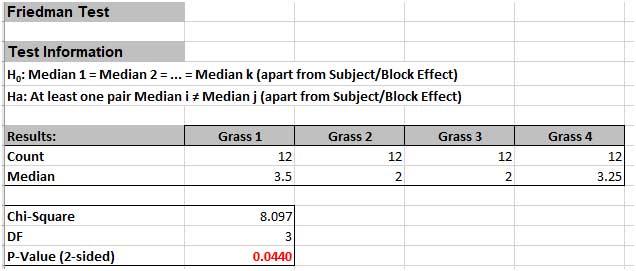
The P-Value of 0.044 tells us that we reject H0. At least one pairwise set of medians are not equal, so there is a difference in the preference of grass types(i.e., there is a difference in treatment effects).
We will now redo the above example to estimate the exact P-Value using Monte Carlo. Typically, this would not be necessary unless the sample sizes were smaller (N subjects <=5 ), but this gives us continuity on the example.

Tip: The Monte Carlo 99% confidence interval for P-Value is not the same as a confidence interval on the test statistic due to data sampling error. The confidence level for the hypothesis test statistic is still 95%, so all reported P-Values less than .05 will be highlighted in red to indicate significance. The 99% Monte Carlo P-Value confidence interval is due to the uncertainty in Monte Carlo sampling, and it becomes smaller as the number of replications increases (irrespective of the data sample size). The Exact P-Value will lie within the stated Monte Carlo confidence interval 99% of the time.
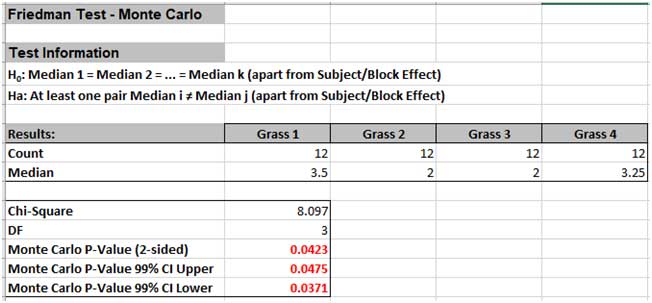
The Monte Carlo P-Value is 0.0475 with a 99% confidence interval of 0.0371 to 0.0475. This will be slightly different every time it is run (the Monte Carlo seed value is derived from the system clock). So, we reject H0: at least one pairwise set of medians are not equal, i.e., there is a difference in the preference of grass types (or there is a difference in treatment effects).
Note that the large sample (asymptotic) P-Value of 0.044 also lies within the Monte Carlo exact confidence interval.